Eigenvalue tracing
I started to work on eigenvalue problems initially for the sake of testing the linear validity of the reduced-dimensional Plesio-Geostrophy (PG) model. Much to my surprise, this seemingly simple and innocent linear problem have been extending endlessly, and I have spent almost \(2/3\) of my doctoral work so far working on eigenvalue problems or normal mode problems in general (not necessarily linked to PG or its successor models).
One of the (many) problems I have encountered in eigenvalue calculations is how to trace an eigenvalue across a parameter space. Suppose we have an eigenvalue-eigenfunction pair \((\lambda_0, \mathbf{x}_0)\) that solves the eigenvalue problem \(\mathbf{A}(\mu) \, \mathbf{x} = \lambda \mathbf{x}\) at \(\mu = \mu_0\), how to get the evolved eigen pair \((\lambda, \mathbf{x})\) at other \(\mu \neq \mu_0\)? In other words, how to calculate (or merely evaluate numerically) the function:
\[\lambda = \lambda(\mu), \quad \mathbf{x} = \mathbf{x} (\mu)\]given their “initial condition”, i.e. the known solution at \(\mu_0\)
\[\lambda(\mu_0) = \lambda_0, \quad \mathbf{x} (\mu_0) = \mathbf{x}_0.\]This problem seems to have a simple solution. We just have to start from \(\mu_0\), take a small step \(\delta \mu\) to \(\mu_1 = \mu_0 + \delta \mu\), and then calculate the eigenvalue of \(\mathbf{A}(\mu_0 + \delta \mu)\) in the vicinity of an initial guess (say \(\lambda_0\)) by introducing shifts and using e.g. the Arnoldi iteration. Following incremental change of \(\mu\), we should, in principle, eventually be able to get to any point \(\mu\) in the parameter space. However, two properties associated with eigen-spectrum distribution pose problems to this straightforward approach.
Eigenvalue crossing / branching / merging
Let us first set up two random matrices, which I will denote \(\mathbf{A}_1\) (left) and \(\mathbf{A}_2\) (right). Each element of these two matrices \(M_{ij} \sim U(-1, 1]\), where \(M=A_{1,2}\):
import numpy as np
rng = np.random.default_rng(42)
dim = 10
A = rng.uniform(-1, 1, size=(dim, dim))
B = rng.uniform(-1, 1, size=(dim, dim))
Their eigenvalues are shown in the complex plane in the middle in blue (\(\mathbf{A}_1\)) and orange (\(\mathbf{A}_2\)), respectively. For general matrices, their eigenvalues will spread all over the complex plane (as shown), with potentially fewer eigenvalues than their dimensions, either due to degeneracy or the matrix being non-normal.
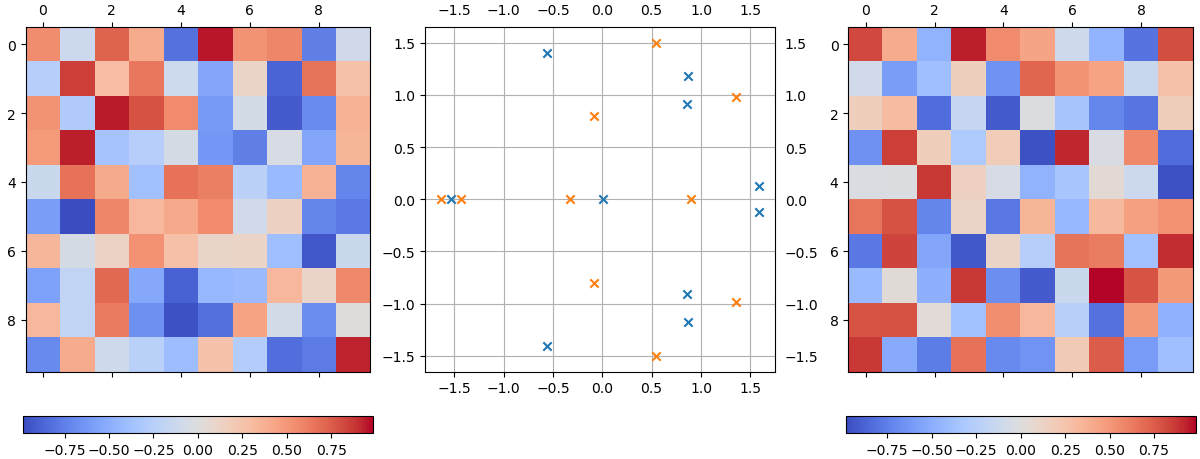
Now let us consider the eigenvalue spectrum of
\[\mathbf{A} = (1 - \mu) \mathbf{A}_1 + \mu \mathbf{A}_2\]The spectrum, \(\lambda\), as a function of \(\mu\), is plotted in three different forms in the figure below.
The bottom left subplot shows the evolution of the eigen-spectrum in the complex plane, with the color labelling the parameter \(\mu\) (color \(\rightarrow\) dark blue when \(\mu \rightarrow 0\), and color \(\rightarrow\) light yellow when \(\mu \rightarrow 1\)). The spectrum evolves from the large dark blue circles representing the spectrum of \(\mathbf{A}_1\), towards the large yellow circles representing the spectrum of \(\mathbf{A}_2\), following the trajectories, as \(\mu\) increases from \(0\) to \(1\). The other subplots show the evolution of \(\mathrm{Re}[\lambda]\) (top left) and \(\mathrm{Im}[\lambda]\) (bottom right) as a function of \(\mu\), respectively.
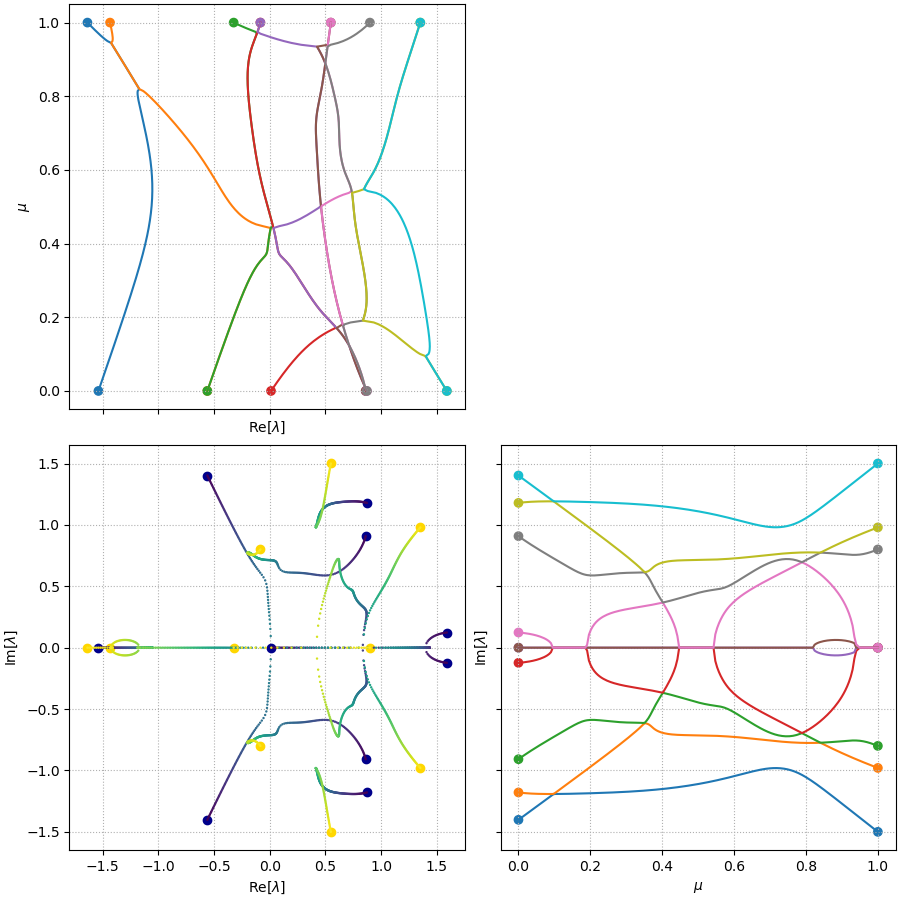
The spectrum evolutions show that there are multiple points where eigenvalues can cross, branch or merge. This phenomenon is more easily seen in the 3D plot of these eigenvalue trajectories (below), where the junctions of trajectories are highlighted. In this random example, we can actually observe (at least) six of such junctions. Three of them (marked by black circles) are junctions where branches visibly merge and branch. Three others (marked by gray circles), are also junctions but just seemingly unconnected due to the finite resolution of the parameter scanning, the sensitive dependency of the eigenvalues on the parameter near the joint, and the saddle-type geometry of the branches.
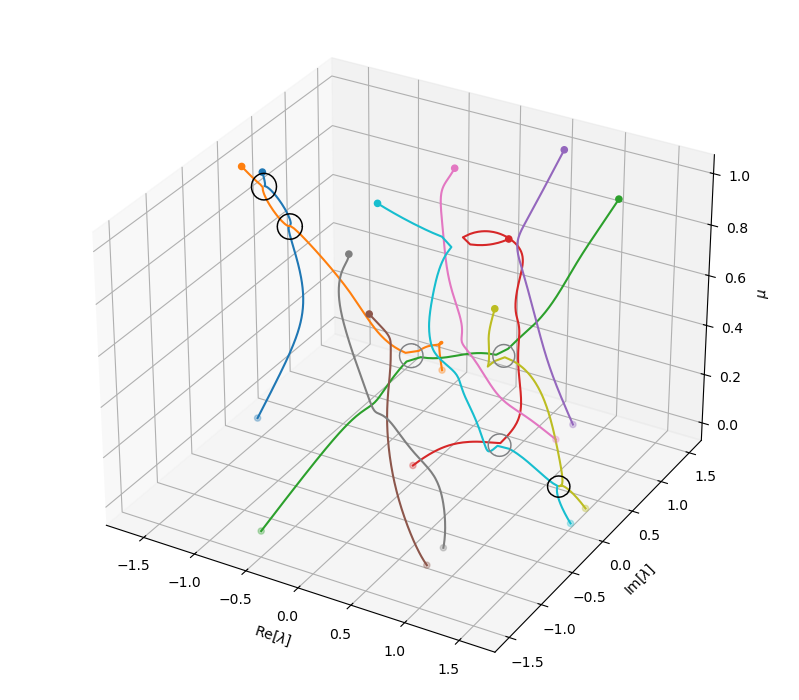
Note that all these visualizations and explorations are only admissible when the system is small enough - in this case, just \(10\times 10\). For high-dimensional systems (which arise, for instance, from discretizing a continuous physical system in high resolutions), there is simply no way to conduct such experiments. Consequently, we have very little prior knowledge of how the eigen-spectrum looks like, or when and where these joints can occur. Tracing an eigenvalue-eigenfunction pair in the parameter space is like holding a bead (the known eigenvalue) on an intangible thread (the “imaginary” branch of this eigenvalue) and looking for the next bead on the same thread in complete darkness (the space of the spectrum). If you take a small step in some direction, you might find your next bead on the same thread, you might find a bead on another irrelevant thread (or many beads on many other threads), or nothing at all - you only know what you get once you make this step and do the calculation, and sometimes a quite expensive one at that.
So what problems do these complicated patterns pose? For starters, these mean that eigenvalues from different branches can get close. If you take a step too far or in the wrong direction, the iterative solver might converge to an eigenvalue that is close to your previous one, but the eigen-pair obtained might belong to a different branch, hence its eigenfunction / eigenvector looks nothing like the previous one. Possible countermeasure to this is to (a) use small enough steps such that the algorithm does not trace to an adjacent branch, and/or (b) wisely predict where the next eigenvalue might be, for instance by extrapolating the previous eigenvalues on the same branch.
Yet another more fundamental problem occurs when branching of an eigenvalue takes place. This usually happens at some degeneracy (or near degeneracy), and as a result, all eigenvalue branches that start from this joint have some level of legitimacy as the successor. In this case, which one do we even trace?
Eigenvalue repulsion / avoided crossing of Hermitian operators
The previous part applies to general square matrices. However, many matrices from physical problems luckily have some structures. Associated with many physical problems such as vibration, diffusion, etc., one important category of matrices / operators for eigenvalue problems is the Hermitian matrices / operators. Such matrices are known to have purely real eigenvalues, as
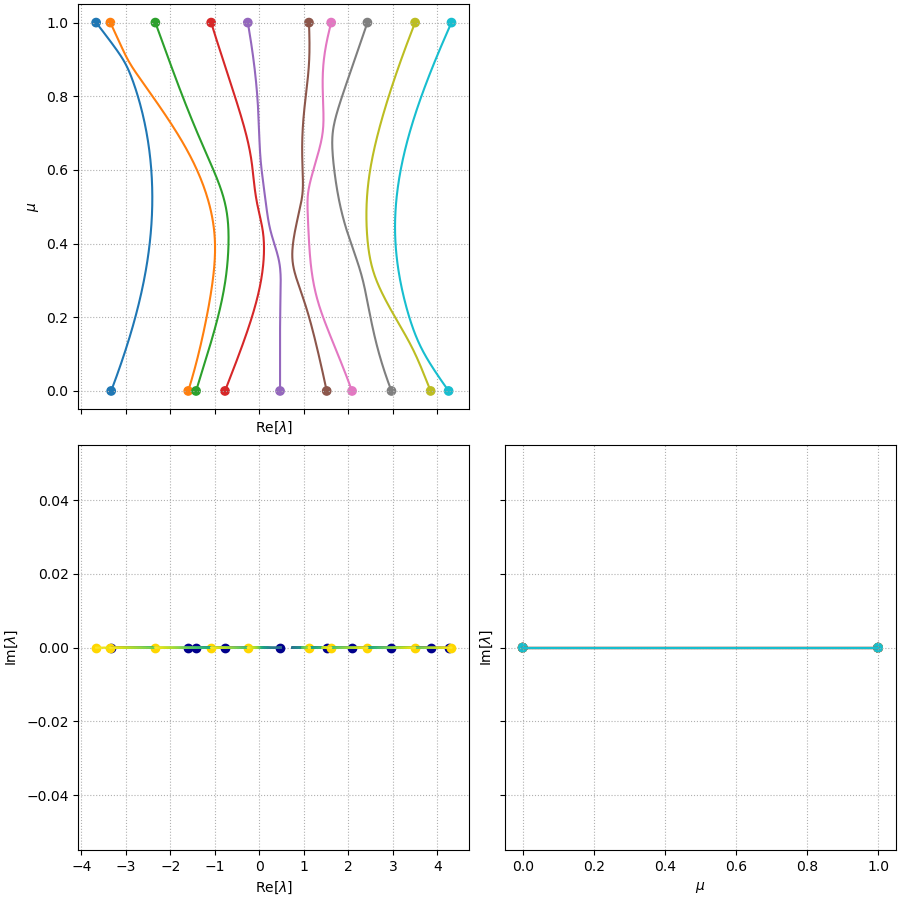
\[\lambda \mathbf{x} = \mathbf{A} \mathbf{x} \quad \Rightarrow \quad \lambda = \frac{\mathbf{x}^H \mathbf{A} \mathbf{x}}{\mathbf{x}^H \mathbf{x}} = \frac{\mathbf{x}^H \mathbf{A}^H \mathbf{x}}{\mathbf{x}^H \mathbf{x}} = \lambda^*.\]The following plot shows two real symmetric matrices, which I will denote \(\mathbf{A}_1\) (left) and \(\mathbf{A}_2\) (right). These are generated from random matrices whose each element \(M_{ij} \sim U(-1, 1]\) and forced to be symmetric via \(\mathbf{A} = \mathbf{M} + \mathbf{M}^T\):
# Force symmetry
A = A.T + A
B = B.T + B
Their eigenvalues are shown in the complex plane in the middle in blue (\(\mathbf{A}_1\)) and orange (\(\mathbf{A}_2\)), respectively.
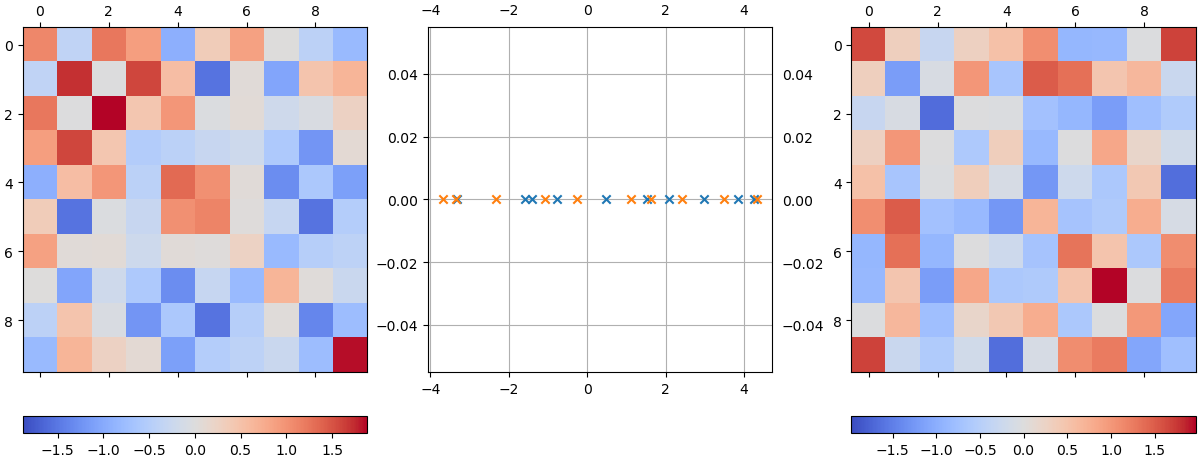
A well-known property of such systems is the eigenvalue repulsion, or avoided crossing. This is the phenomenon that as two eigenvalues \(\lambda_1\), \(\lambda_2\) evolve with \(\mu\), their trajectories will often deflect and get repelled from one another when they become too close.
As an example, I have calculated the eigenvalue spectrum of
\[\mathbf{A} = (1 - \mu) \mathbf{A}_1 + \mu \mathbf{A}_2\]for an array of \(\mu\). The \(\lambda(\mu)\) curves calculated this way are shown in the following figure. Clear deflection can be seen for the orange / blue lines (corresponding to \(\lambda_1\), \(\lambda_2\) given \(\lambda_1 \leq \lambda_2 \leq \dots \lambda_{10}\)) at \(\mu \approx 0.9\), purple / brown lines (\(\lambda_5\) and \(\lambda_6\)) at \(\mu\approx 0.35\), and pink / brown / gray lines (\(\lambda_6\), \(\lambda_7\) and \(\lambda_8\)) at \(\mu\approx 0.5, 0.7\), etc.
The phenomenon of eigenvalue repulsion is apparently of research interest themselves. For the eigenvalue tracing problem per se, this phenomenon has both its upside and downside. As an upside, it actually counteracts the crossing, merging and branching of general matrices, hence its alternative name “avoided crossings”. This means that different branches of eigenvalues would typically have some distance apart, which would help to prevent eigenvalue tracing algorithm from erroneously jumping between branches.
However, the phenomenon also means that eigenvalue branches would be deflected in the complex plane, taking shapes that they otherwise wouldn’t take if not for an adjacent eigenvalue to repel them. Consequently, sophisticated predictors of the eigenvalue evolution might predict a trend of \(d\lambda/d\mu\) inconsistent with the deflected trend. Applying such predictors might actually make it more possible for the eigenvalue tracing algorithm to jump to the adjacent branch.
It is also worth noticing that most of real world applications are somewhere in between: they are described not by purely Hermitian operators, but also not by purely random non-Hermitian operators. A damped harmonic oscillator, described by
\[\frac{d^2 x}{dt^2} + c \frac{dx}{dt} + x = 0\]for instance, when introducing \(y=dx/dt\), can be described as a first-order dynamical system by a skew-symmetric operator, with a weak diagonal operator proportional to the damping:
\[\frac{d}{dt} \begin{pmatrix} x \\ y \end{pmatrix} = \begin{pmatrix} 0 & 1 \\ -1 & -c \end{pmatrix} \begin{pmatrix} x \\ y \end{pmatrix}.\]Similar structure applies to any Hamiltonian system with additional damping. In addition, many nonlinear systems, when linearized around a non-zero background field, are in general non-Hermitian, but might contain a large Hermitian part. For all these problems, all the phenomena described above, including eigenvalue merging and branching, crossing and avoided crossing, can occur in the same system. Reusing our analog of bead-searching, while searching for the next bead in the dark, we need to worry about:
- If we find another bead by randomly searching in the vicinity of the current bead, could it be that this bead belongs to a different thread? (\(\rightarrow\) eigenvalue crossing / close-by branch)
- If we guide our search by taking a step in the direction formed by the previous beads, could it be that we miss the next beam since the thread has already changed its course, possibly deflected by another thread nearby? (\(\rightarrow\) eigenvalue repulsion / avoided crossing)
- If we find multiple beads in the vicinity of our prediction that all resemble the current bead to some extent, how should we decide which one is the next one on the thread? Or has the thread itself branched into multiple threads? (\(\rightarrow\) branching)
Algorithmic solutions
There are some methods / algorithms that, I believe, can counteract the problems listed above. All these algorithms are amendments to a vanilla version of the eigenvalue tracing procedure, described by the following pseudocode:
Given mu0, mu1, dmu, ev0
mu_array = range(start=mu0, stop=mu1, step=dmu)
ev_array = array(shape=(mu_array.shape))
ev_array[0] = ev0
for i in range(start=1, stop=mu_array.size):
mu = mu_array[i]
A = get_operator(parameter=mu)
ev_tmp = solve_eigenvalue(matrix=A, target=ev_array[i-1])
ev_array[i] = ev_tmp
Accurate predictor
Most iterative solvers for sparse systems work best if it has a starting guess of the eigenvalue it is seeking. In the vanilla algorithm pseudocode, we are supplying just the previous eigenvalue, ev_array[i-1]. While it may suffice in many cases, it would be problematic when a branch crossing occurs. If two branches non-tangentially cross each other, algorithms using the previous eigenvalue as the starting guess would likely be sucked to the crossing branch instead of the original branch. In this case of branch crossing, a better predictor can be provided such that the iterative algorithm has better chance to converge to the desired eigenvalue.
One option is to use extrapolation to better predict where the next eigenvalue lies. The simplest would be a linear extrapolation
\[\hat{\lambda}_i = \frac{\mu_i - \mu_{i-2}}{\mu_{i-1} - \mu_{i-2}} \lambda_{i-1} + \frac{\mu_i - \mu_{i-1}}{\mu_{i-2} - \mu_{i-1}} \lambda_{i-1}\]hence motivating the amendment
ev_predict = (mu_array[i] - mu_array[i-2])/(mu_array[i-1] - mu_array[i-2])*ev_array[i-1]
+ (mu_array[i] - mu_array[i-1])/(mu_array[i-2] - mu_array[i-1])*ev_array[i-2]
ev_tmp = solve_eigenvalue(matrix=A, target=ev_predict)
Other more complicated extrapolation schemes can of course be swapped in the predictor. We just have to note that extrapolation schemes only work when several eigenvalues in this branch have already been computed such that \(i-1\), \(i-2\) (and other previous eigenvalues used in the extrapolation) are well defined.
Another issue with these extrapolation-based predictors is that since they are often built on polynomials, they are quite bad at capturing power law depedencies. In many cases I work with, \(\Delta \lambda \sim \Delta \mu^\alpha\) with \(\alpha\) potentially \(\neq 1\), which will not be accurately predicted by the extrapolation. Nevertheless, simple schemes such as linear extrapolation will mostly predict the correct linear trend, and can at least be useful in the case of oblique eigenvalue crossings. Therefore I routinely use linear extrapolation for tracing the next eigenvalue (I have also used higher-order extrapolations but I haven’t yet seen their advantage over linear extrapolations).
The second option, which I also routinely apply, is to supply not only an eigenvalue seed but also an initial guess on the eigenvector. The revised pseudocode would then read
evec_array = array(...)
for i in range(start=1, stop=mu_array.size):
...
ev_tmp, evec_tmp = solve_eigenvalue(matrix=A, target=ev_predict, v0=evec_array[i-1])
ev_array[i] = ev_tmp
evec_array[i] = evec_tmp
This requires the additional cost to at least store one eigenvector along the way.
Similarity-based filtering
This is a simple countermeasure to the risk of jumping to the wrong branch. Once a new eigenvalue is obtained, we verify that it belongs to the branch currently being traced by checking if it is similar enough to our expectation. Typically, this can comprise of two criteria:
- The eigenvalue criterion: the newly obtained eigenvalue \(\lambda_i\) should fall close enough to the initial prediction \(\hat{\lambda}_i\):
where \(\epsilon_1\) and \(\epsilon_2\) are small numbers for thresholds. \(\epsilon_2\) is useful when one of the eigenvalues are too close to zero.
- The eigenvector criterion (optional, depending on if eigenvector is stored): the newly obtained eigenvector \(\mathbf{x}_i\) should be similar enough to the previous one, i.e.
One can show this metric, similar to \(1 - R^2\), is the smallest misfit between the two vectors under arbitrary rotation.
Collecting several candidate solutions from the eigensolver and filtering using these criteria guarantees that the eigen-pairs left are at least similar enough to the previous one on the branch, and has legitimacy of being the next in line. For systems forced to be symmetric / Hermitian or skew-symmetric, it usually works well since the eigenvectors of different eigenvalues are orthogonal to one another, hence the eigenvalue criterion serves as a powerful tool to filter out those on different branches. However, this method will have its problem in case of: (1) degeneracy (which should be rare in a transitional state), (2) eigenvalue crossing for non-Hermitian systems (in which case eigenvectors can also be similar).
While not sufficient to pick out the correct ones in all scenarios, this method nevertheless provides necessary conditions for tracing eigen-pairs, hence is almost always implemented in my code. The only time I forgot about this amendment, the result was… complicated.
Adaptive parameter step
To be developed…
Branched tracing
So far none of the amendments can tackle the situation where the eigenvalue just branches into several eigenvalues. It seems to me that there are simply no shortcut without further prior knowledge, but to either randomly choose one of the branched eigenvalues, or to trace all simultaneously.
By implementing this on the aforementioned toy example of random matrices, I show that simultaneous tracking several branches is possible (see plot below for the result of eigenvalue tracing using the second eigenvalue of \(A_1\) as the starting point at \(\mu=0\)).
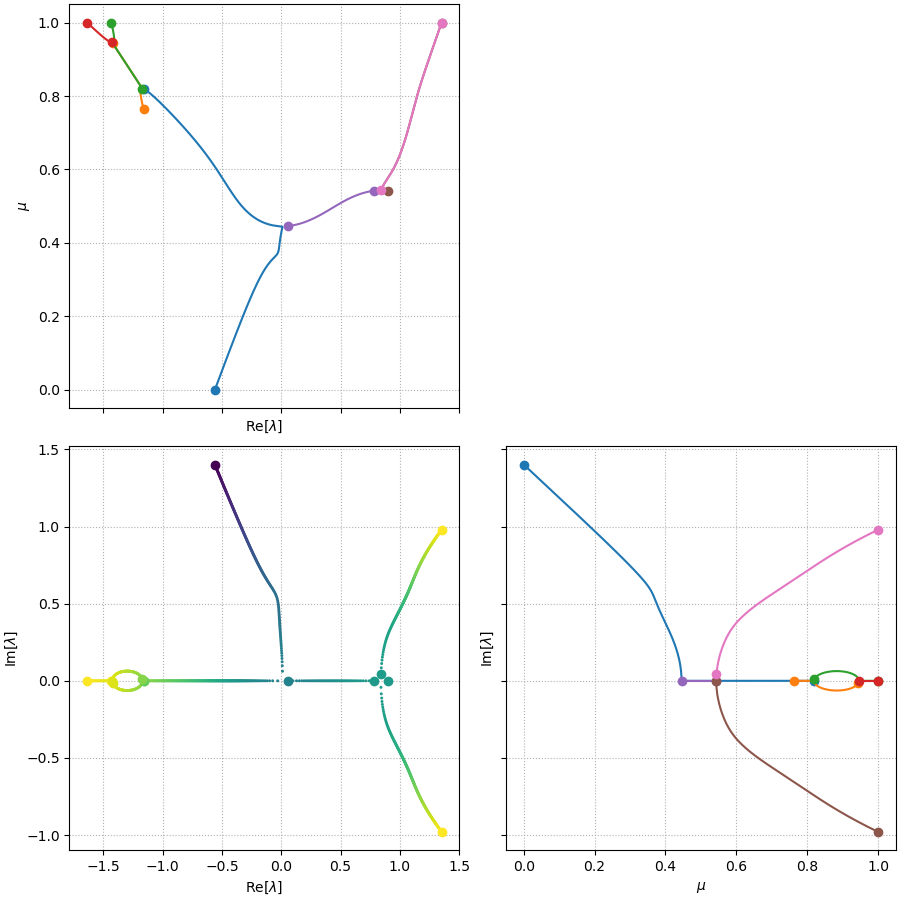
In fact, it actually provides a much enriched picture of the otherwise darker space of eigen-spectrum, also illustrated by the 3D branch plot below.
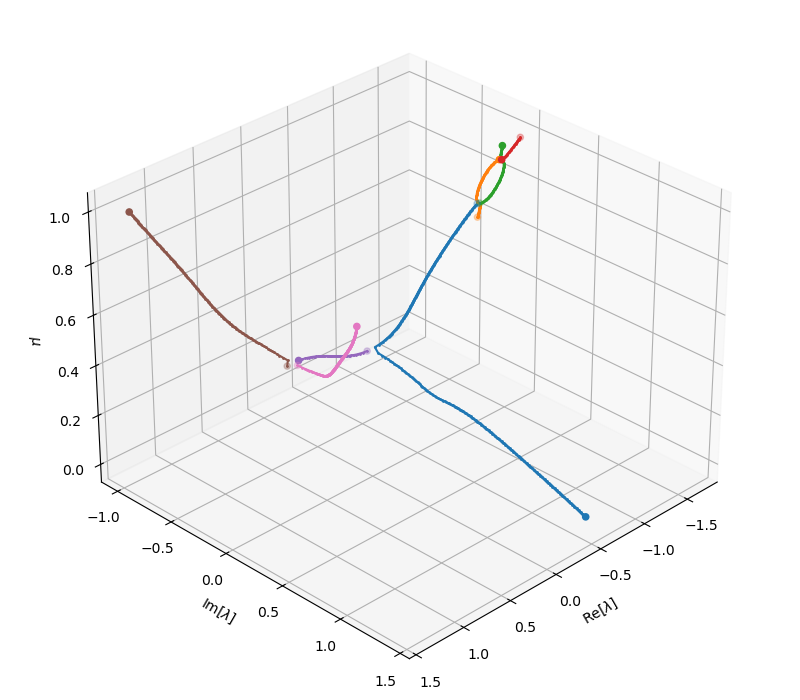
Tracing multiple branches should only be used in combination with the aforementioned similarity-based filtering. This is because the eigensolver can almost always return several eigen-pairs. Proceeding to trace them all without verifying their validity will result in exponential growth of number of traced branches, which will be computationally completely prohibitive.
Another subtlety of this algorithm is when branching occurs, the algorithm will naturally discover several viable eigen-pair options and start tracing them simultaneously. At the next parameter step, since the multiple branches are still very close together, each branch will still discover the eigenvalues on adjacent branches. If one naively proceeds to trace them all, one will again have exponentially growing number of branches due to repeated registration of the same branches.